Our immune system relies on T cells to fight infections. Like a sports team, T cells don’t just show up and react — they also rely on the lymphoid organs as their home base, where they train, get their game plan, and coordinate their defenses.
Researchers have struggled to understand how this counteroffensive evolves across these sites. Now, a new tool from Biohub allows scientists to permanently tag recently activated T cells with a fluorescent protein to track how they travel and change during an infection. The system, recently described in Nature Immunology, makes it possible to precisely characterize the T cells that respond to a specific threat and understand how their location shapes them.
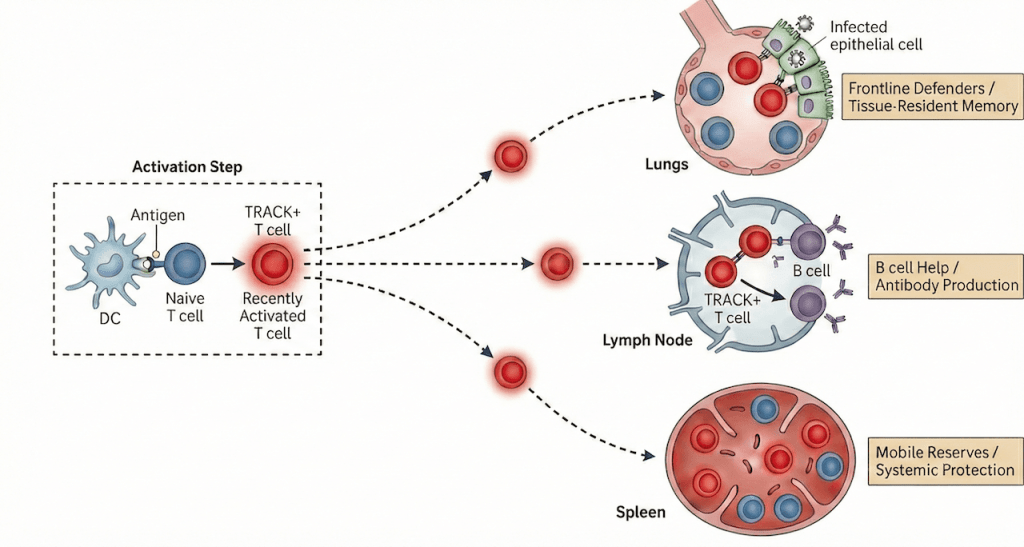
Researchers from Biohub and The Rockefeller University used the tool, called TRACK (Tracking Recently Activated Cell Kinetics), to study T cells across three sites — the lungs, lymph nodes, and spleen — as they mounted and later maintained a defense against the flu virus. Their data revealed a “division of labor” among these locations.
The study is a critical step in pushing Biohub towards one of its core science goals – reprogramming the immune system to better respond to aging and disease. “To reprogram the immune system, we first have to understand the genetic programs that guide immune cells across different organs,” said first author Roham Parsa, group leader of Immune Cell Dynamics and Function at Biohub and adjunct assistant professor at The Rockefeller University. “By mapping how these programs change from tissue to tissue, we can begin to design strategies to precisely redirect immune function.”
He added: “The T cell response is compartmentalized in a very elegant way.” Parsa’s group aims to understand how immune cells adapt, communicate, and redistribute across organs over time, uncovering the cellular dynamics that shape protective immunity.
He and the team found that the tissue where T cells first encounter the virus shapes their identity: Lung T cells become specialized frontline defenders, lymph node T cells focus on helping B cells make antibodies, and spleen T cells act as mobile reserves that spread protection across the body. Over time, these populations redistribute and converge into a shared immune memory, equipping the body for faster responses in the future.
Although this study focused on flu infection, the TRACK system — which is genetically bioengineered into mice — could be applied beyond infection to study cancer, vaccination, and autoimmune conditions.
The call to action
After the immune system detects something foreign, such as viral particles infiltrating the lungs, it formulates a precision attack. This adaptive response primes defenses to recognize the enemy based on bits of its proteins, called antigens. T cells become activated after encountering antigens displayed by other immune cells called antigen-presenting cells.
Newly mobilized T cells multiply, forming genetically identical lineages called clones that either directly attack the pathogen or coordinate with other immune cells to do so. After the danger has passed, a portion of these cells remain, serving as long-lived memory T cells that protect against future infections.
Previous studies suggested that T cell responses differ between infected tissue and lymphoid tissue, including lymph nodes and spleen. But it has been difficult to capture these variations as an immune response unfolds.
“If you want to study cells responding to a particular infection, a big problem is, where are you going to start?” said senior author Daniel Mucida, an Affiliated Investigator at Biohub and professor at The Rockefeller University. “Because there are so many T cells already present, you can’t distinguish only the cells that interest you.”
Spotting the right T cells
Parsa addressed this challenge by focusing on a fleeting combination of proteins expressed together briefly after a T cell encounters an antigen. He built the TRACK system so that when a T cell in a mouse expresses these two molecules at the same time, a glowing red label switches on. This signal allowed researchers to pinpoint the T cells responding to that antigen and follow them over time, tracking their movement between organs and how their roles evolved.
In their experiments, the researchers studied cells known as CD4+ T cells. Once newly activated CD4+ T cells were labeled, they analyzed them by sequencing their RNA and the genetic code of their antigen receptors. They examined these cells nine days after infection, when the immune response was at its peak, and again, long after the response had subsided. Differences among the three organ sites revealed an evolving, and likely strategic, response.
Flexible cells, evolving response
As the primary site of flu infection, the lungs shaped local T cell identity. More than in other organs, the lungs contained CD4+ T cells capable of directly destroying infected host cells. Weeks later, many of these lung T cells became tissue-resident memory cells.
In contrast, the spleen functioned as a supply depot, contributing a substantial portion of T cells to the lungs during infection. The lymph nodes contained T cells primarily focused on helping B cells produce antibodies.
The data also revealed T cell flexibility. Depending on site and timing, genetically identical clones fulfilled different roles — assisting antibody production, killing infected cells, or providing long-term protection.
“Even though they are responding to the same pathogen, they perform different functions that are involved in different aspects of adaptive immunity,” Mucida said.
The distribution of T cell clones across organs reflected the antigens they targeted — a legacy of initial priming. However, compartmentalization was not absolute. T cells reacting to different viral proteins appeared in multiple locations, suggesting tissues do not strictly segregate responses. Instead, each site favors expansion of certain clones, likely because distinct tissues display different viral fragments that selectively stimulate specific T cells.
“It’s like athletes who could play for many teams, but each team emphasizes different positions, so players who best fit that strategy get more time on the field and gradually make up a larger share of the lineup,” Parsa explained.
Over time, these differences became less pronounced as clones migrated, particularly from the lungs to the lymph nodes, leading to a more even distribution.“You can speculate about what this means,” Parsa said. “I think it’s a way of preparing T cells in case the pathogen is reintroduced in a different location.”
Reprogramming the immune response
The researchers are now applying TRACK to other diseases. Mucida has begun studying T cell responses in early-stage colorectal cancer and how tumors evade them. Parsa is examining autoimmune and neurodegenerative diseases such as multiple sclerosis, in which misguided T cells damage neurons.
A more detailed understanding of immune dynamics could lead to improved strategies for manipulating immunity. Parsa aims to uncover how T cells communicate with tissues and how local environments shape their function. By identifying these tissue-specific cues, researchers may learn how to guide immune responses more precisely in different organs and diseases.
He expects immune dynamics to differ across conditions, and envisions the TRACK mice as a tool to help reveal those patterns. “My hope is that it will allow other scientists to look at new pathogens and other perturbations more efficiently,” he said.