Synthetic Spatial Omics
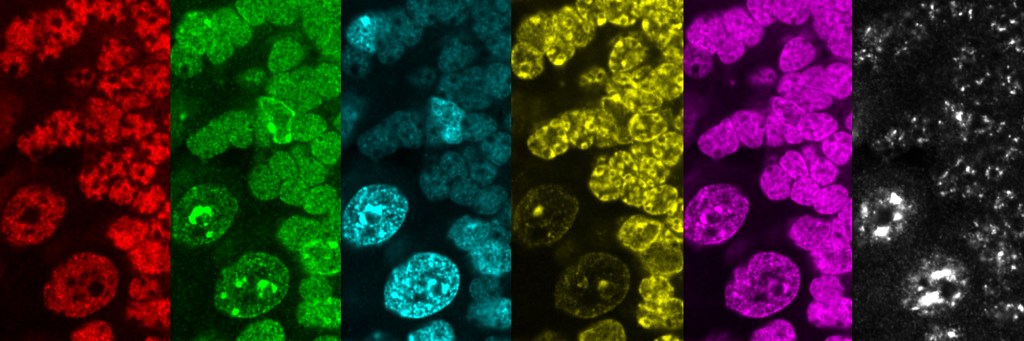
Decoding and engineering cell state dynamics through spatial organization
How cells process signals and make decisions depends fundamentally on the spatial organization of molecules within cells and how this organization changes over time. To decode these mechanisms and enable rational cell engineering, we develop technologies that measure, perturb, and control molecular states in single cells. By integrating synthetic biology, spatial multi-omics, systems biology, and AI-driven approaches, we aim to understand how molecular architecture drives cell state transitions and to leverage this knowledge to program cellular behaviors for therapeutic applications.
Research
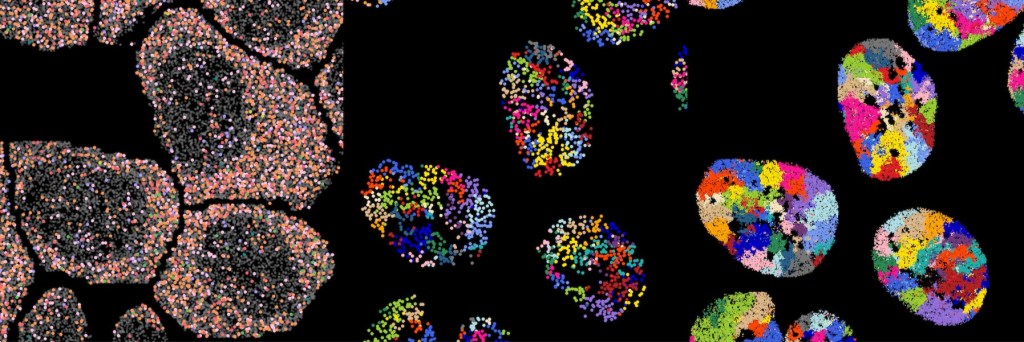
Decoding molecular architecture during cell state transitions
Cell states emerge from the spatial organization and interactions of proteins, nucleic acids, and other biomolecules. To understand how cells interpret complex external signals through these spatial arrangements, we develop scalable multi-omics platforms that measure subcellular reorganization across combinatorial perturbations. By profiling how molecular neighborhoods change in response to defined inputs over time, we identify organizational principles that govern gene regulation and cellular decision making.
Engineering programmable cell functions with spatial precision
Cell engineering would benefit from understanding where disease signals emerge in tissues, when cells change state, and how therapeutic payloads reshape cellular environments. We engineer cells with synthetic biology toolkits, then use spatial profiling to decode how these engineered systems interact with their molecular neighborhoods. This approach accelerates therapeutic cell engineering while revealing fundamental principles of spatial organization.
Developing next-generation spatiotemporal genomics technologies
Understanding how past signals and molecular events influence future cell states requires measuring gene regulation across both space and time. We develop molecular recording technologies that write cellular history into the cells, which can be read out alongside spatial multi-omics measurements. These approaches reveal how earlier molecular events bias cell fate decisions, ranging from development to disease progression, and identify causal molecular determinants of cell state transitions.